Overview
This article uses the public survival::veteran dataset
as a compact real-data example. It is not a production oncology
registry, but it provides a real patient-level time-to-event dataset
with treatment assignment and baseline covariates.
Prepare the public dataset
library(survival)
data(veteran, package = "survival")
#> Warning in data(veteran, package = "survival"): data set 'veteran' not found
veteran2 <- veteran
veteran2$arm <- ifelse(veteran2$trt == 1, "SOC", "A")
veteran2$event <- veteran2$status
veteran2$subgroup <- as.character(veteran2$celltype)
veteran2$prior <- factor(veteran2$prior)
head(veteran2)
#> trt celltype time status karno diagtime age prior arm event subgroup
#> 1 1 squamous 72 1 60 7 69 0 SOC 1 squamous
#> 2 1 squamous 411 1 70 5 64 10 SOC 1 squamous
#> 3 1 squamous 228 1 60 3 38 0 SOC 1 squamous
#> 4 1 squamous 126 1 60 9 63 10 SOC 1 squamous
#> 5 1 squamous 118 1 70 11 65 10 SOC 1 squamous
#> 6 1 squamous 10 1 20 5 49 0 SOC 1 squamous
profile_cce_dataset(
data = veteran2,
arm = "arm",
time = "time",
event = "event",
subgroup = "subgroup"
)
#> CCE dataset profile
#> n events event_rate median_follow_up max_follow_up
#> 1 137 128 0.9343066 80 999Fit a VS comparison
vs_fit <- fit_cce_vs(
data = veteran2,
arm = "arm",
time = "time",
event = "event",
covariates = c("karno", "diagtime", "age", "prior", "subgroup"),
subgroup = "subgroup",
tau = 180,
landmark_times = c(90, 180),
bootstrap = 10,
seed = 300
)
summary(vs_fit)
#> CCE VS result
#> Label: ok
#> Warnings: Residual covariate imbalance detected.
#> mode method subgroup tau rmst_arm0 rmst_arm1 delta_rmst landmark_time
#> 1 vs gformula All 180 17.42348 13.70885 -3.714629360 90
#> 2 vs gformula All 180 17.42348 13.70885 -3.714629360 180
#> 3 vs iptw_km All 180 94.46028 81.83579 -12.624490041 90
#> 4 vs iptw_km All 180 94.46028 81.83579 -12.624490041 180
#> 5 vs iptw_cox All 180 88.51387 88.51779 0.003925953 90
#> 6 vs iptw_cox All 180 88.51387 88.51779 0.003925953 180
#> survival_arm0 survival_arm1 delta_survival delta_rmst_lower_ci
#> 1 0.022844751 0.0106242942 -1.222046e-02 -9.131745
#> 2 0.001166565 0.0002869724 -8.795925e-04 -9.131745
#> 3 0.544475074 0.4012334748 -1.432416e-01 -30.010487
#> 4 0.191913854 0.2287792627 3.686541e-02 -30.010487
#> 5 0.475309651 0.4753349641 2.531322e-05 -11.082825
#> 6 0.212142801 0.2121663536 2.355226e-05 -11.082825
#> delta_rmst_upper_ci delta_survival_lower_ci delta_survival_upper_ci
#> 1 -0.9845419 -0.039905898 -1.360854e-04
#> 2 -0.9845419 -0.024627849 -1.241037e-09
#> 3 -4.4122734 -0.272187557 -7.639318e-02
#> 4 -4.4122734 -0.006723102 9.796009e-02
#> 5 7.6966722 -0.072056221 4.926232e-02
#> 6 7.6966722 -0.062511997 4.403169e-02
plot(vs_fit, method = "iptw_km", subgroup = "All")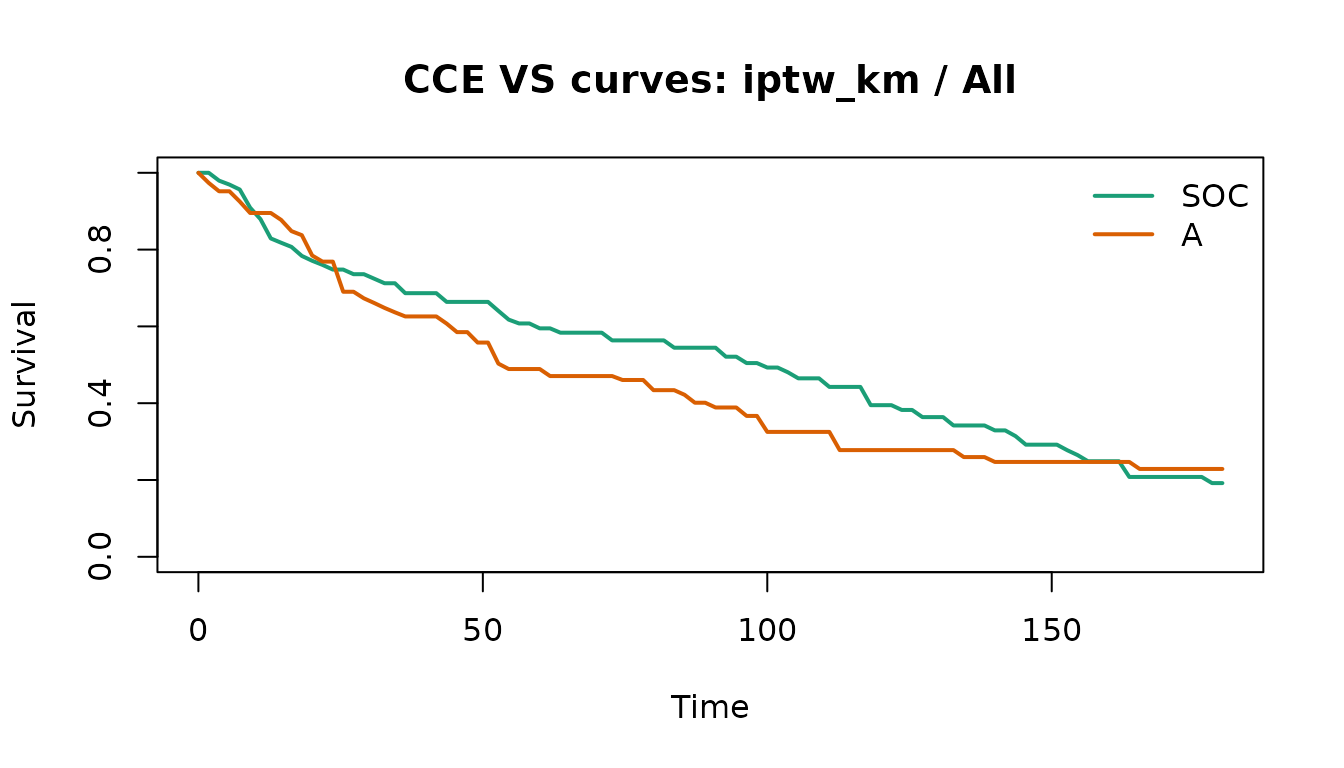
head(as_effects_df(vs_fit))
#> mode method subgroup tau rmst_arm0 rmst_arm1 delta_rmst landmark_time
#> 1 vs gformula All 180 17.42348 13.70885 -3.714629360 90
#> 2 vs gformula All 180 17.42348 13.70885 -3.714629360 180
#> 3 vs iptw_km All 180 94.46028 81.83579 -12.624490041 90
#> 4 vs iptw_km All 180 94.46028 81.83579 -12.624490041 180
#> 5 vs iptw_cox All 180 88.51387 88.51779 0.003925953 90
#> 6 vs iptw_cox All 180 88.51387 88.51779 0.003925953 180
#> survival_arm0 survival_arm1 delta_survival delta_rmst_lower_ci
#> 1 0.022844751 0.0106242942 -1.222046e-02 -9.131745
#> 2 0.001166565 0.0002869724 -8.795925e-04 -9.131745
#> 3 0.544475074 0.4012334748 -1.432416e-01 -30.010487
#> 4 0.191913854 0.2287792627 3.686541e-02 -30.010487
#> 5 0.475309651 0.4753349641 2.531322e-05 -11.082825
#> 6 0.212142801 0.2121663536 2.355226e-05 -11.082825
#> delta_rmst_upper_ci delta_survival_lower_ci delta_survival_upper_ci
#> 1 -0.9845419 -0.039905898 -1.360854e-04
#> 2 -0.9845419 -0.024627849 -1.241037e-09
#> 3 -4.4122734 -0.272187557 -7.639318e-02
#> 4 -4.4122734 -0.006723102 9.796009e-02
#> 5 7.6966722 -0.072056221 4.926232e-02
#> 6 7.6966722 -0.062511997 4.403169e-02Create an SOC-only projection
For scenario planning, we can keep only the SOC rows and overlay assumption-based proportional-hazards projections.
soc_fit <- project_soc_only(
data = veteran2,
arm = "arm",
soc_level = "SOC",
time = "time",
event = "event",
subgroup = "subgroup",
tau = 180,
hr_scenarios = c(0.75, 0.90, 1.00),
target_delta_rmst = 15,
prior_mean_log_hr = log(0.85),
prior_sd_log_hr = 0.20,
bootstrap = 10,
seed = 400
)
summary(soc_fit)
#> CCE SOC-only projection
#> Label: Projection (assumption-based)
#> mode method subgroup scenario_hr tau rmst_arm0 rmst_arm1
#> 1 soc_only projection_ph All 0.75 180 96.12878 110.83745
#> 2 soc_only projection_ph All 0.75 180 96.12878 110.83745
#> 3 soc_only projection_ph All 0.90 180 96.12878 101.65283
#> 4 soc_only projection_ph All 0.90 180 96.12878 101.65283
#> 5 soc_only projection_ph All 1.00 180 96.12878 96.12878
#> 6 soc_only projection_ph All 1.00 180 96.12878 96.12878
#> delta_rmst landmark_time survival_arm0 survival_arm1 delta_survival
#> 1 14.708678 90 0.5467462 0.6358276 0.08908133
#> 2 14.708678 180 0.2124268 0.3129010 0.10047422
#> 3 5.524058 90 0.5467462 0.5807741 0.03402784
#> 4 5.524058 180 0.2124268 0.2480209 0.03559415
#> 5 0.000000 90 0.5467462 0.5467462 0.00000000
#> 6 0.000000 180 0.2124268 0.2124268 0.00000000
#> required_hr pos_proxy delta_rmst_lower_ci delta_rmst_upper_ci
#> 1 0.7455392 0.258 13.485943 15.473156
#> 2 0.7455392 0.258 13.485943 15.473156
#> 3 0.7455392 0.258 5.116966 5.797128
#> 4 0.7455392 0.258 5.116966 5.797128
#> 5 0.7455392 0.258 0.000000 0.000000
#> 6 0.7455392 0.258 0.000000 0.000000
#> delta_survival_lower_ci delta_survival_upper_ci
#> 1 0.07880162 0.09380862
#> 2 0.08812526 0.10462482
#> 3 0.03039300 0.03562514
#> 4 0.03010239 0.03781181
#> 5 0.00000000 0.00000000
#> 6 0.00000000 0.00000000
plot(soc_fit, subgroup = "All")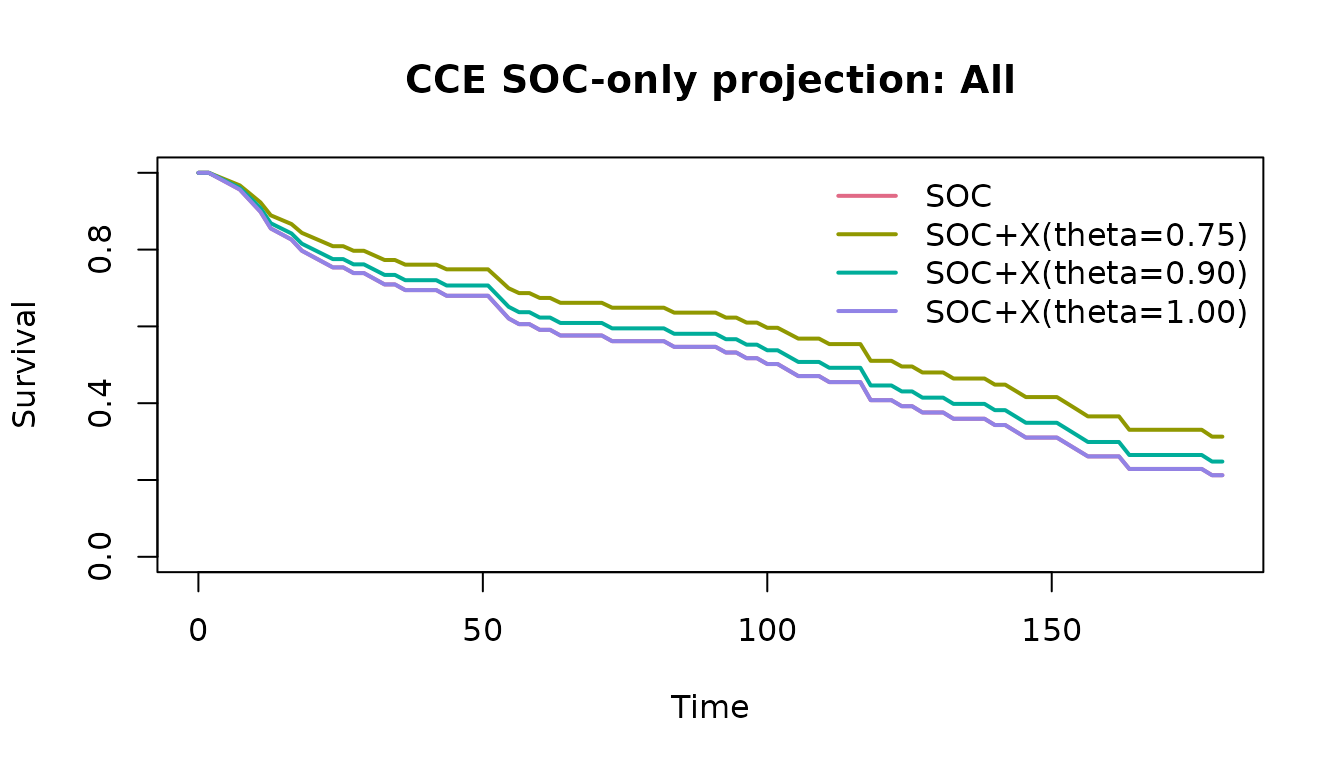
Interpretation notes
- VS-mode results include
gformula,iptw_km, andiptw_cox, so the method label should be interpreted explicitly. - SOC-only outputs are projections, not causal estimates.
- In real programs, diagnostics should be interpreted together with data provenance, treatment policy definitions, and endpoint adjudication rules.